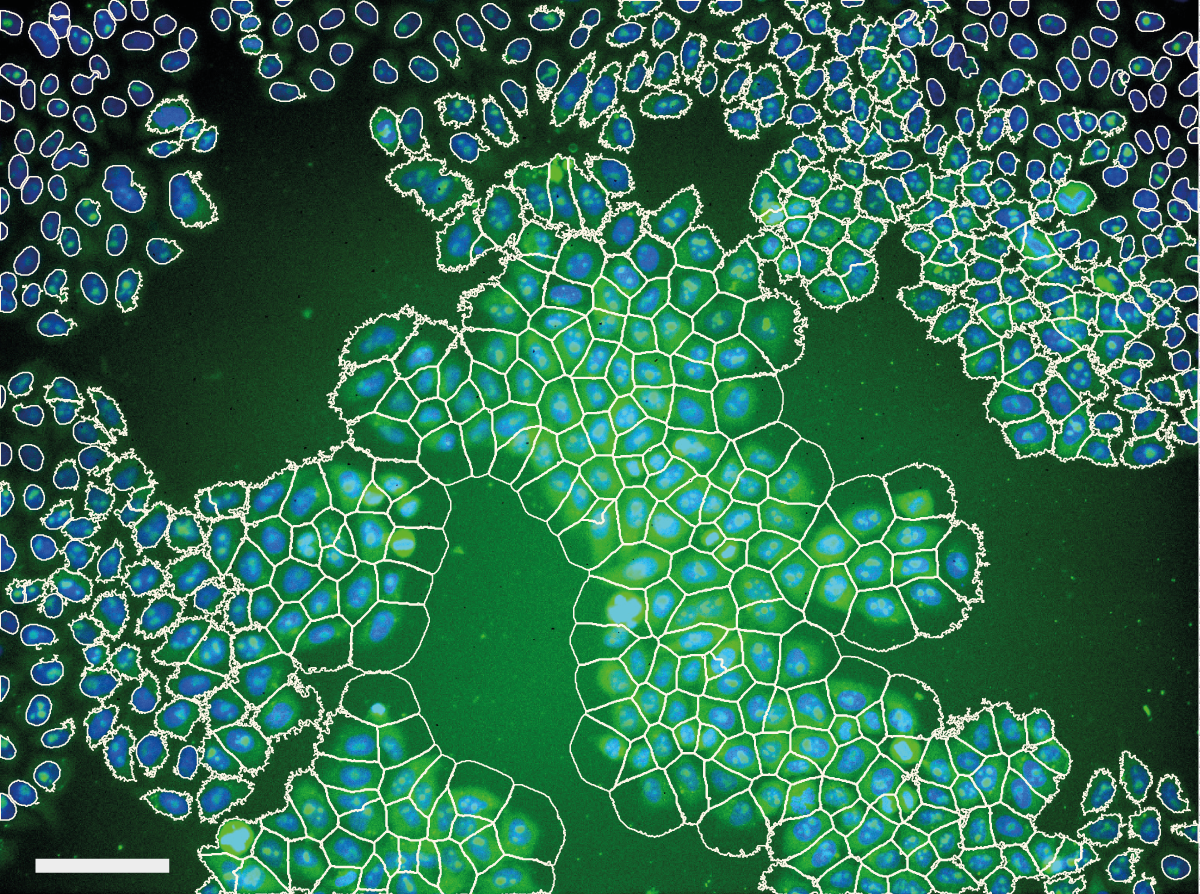
 
They are also able to deal with both two-dimensional and three-dimensional (3D) information, thus laying the foundations for comprehensive molecular maps of tissues and organs. Both data representations leverage sparse 18 or memory-efficient 19 approaches in Python for scalability and ease of use. 
Squidpy introduces two main data representations to manage and store spatial omics data in a technology-agnostic way: a neighborhood graph from spatial coordinates and large-source tissue images acquired in spatial omics data (Fig. 
Squidpy aims to bring the diversity of spatial data in a common data representation and provide a common set of analysis and interactive visualization tools. A comprehensive framework that enables community-driven scalable analyses of both spatial neighborhood graph and image, along with an interactive visualization module, is missing (Supplementary Table 1).įor this purpose we developed ‘Spatial Quantification of Molecular Data in Python’ (Squidpy), a Python-based framework for the analysis of spatially resolved omics data (Fig. The combination of different analysis steps is still hampered by the lack of a unified data representation and of a modular application programming interface, for example loading processed data from Starfish 5, combining stLearn’s 11 integrative analysis of tissue images together with Giotto’s powerful spatial statistics 13, BayesSpace spatial clustering 14 or leveraging state-of-the-art deep-learning-based methods for image segmentation 15, 16 and visualization 17. The underlying computational challenges lie in efficient data representation as well as comprehensive analysis and visualization methods.Įxisting analysis frameworks for spatial data focus either on preprocessing 5, 6, 7, 8 or on one particular aspect of spatial data analysis 9, 10, 11, 12, 13. Such diversity in generated data and corresponding formats currently represents an infrastructural hurdle that has hampered urgently needed development of interoperable analysis methods. In contrast to the current state-of-the-art dissociation-based protocols, spatial molecular technologies acquire data in greatly diverse forms, in terms of resolution (few cells per observation to subcellular resolution), multiplexing (dozens of features to genome-wide expression profiles), modality (transcriptomics, proteomics and metabolomics) and often with an associated high-content image of the captured tissue 2, 3, 4. Spatially resolved molecular technologies aim at bridging this gap by enabling the investigation of tissues in situ at cellular and subcellular resolution 2, 3, 4. However, how cellular diversity constitutes tissue organization and function is still an open question. Dissociation-based single-cell technologies have enabled the deep characterization of cellular states and the creation of cell atlases of many organs and species 1. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
